This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
Introduction
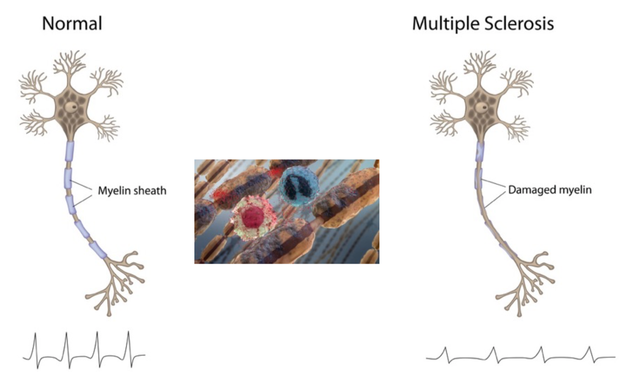
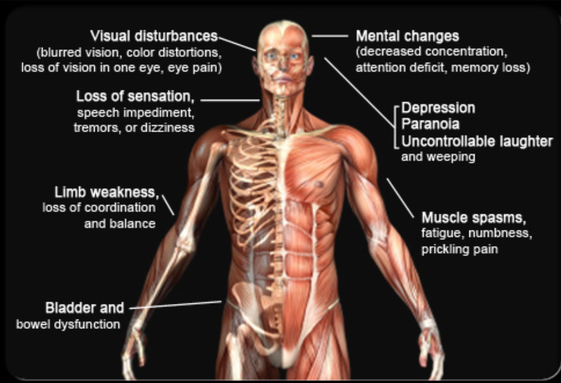
Multiple sclerosis is an immune-mediated disease that affects the brain and spinal cord. MS is considered to be immune-mediated instead of autoimmune because the exact target of the immune cells is unknown. In this disease, the body’s own immune system attacks and degrades the myelin sheath that protect nerve fibers. This causes communication between the brain and the rest of the body to be hindered [1]. As the disease progresses the nerves themselves can begin to degrade and permanently damaged. The symptoms can be varied due to the signals sent by the brain through the nerves becoming distorted. MS has four disease courses with varying severities [2].
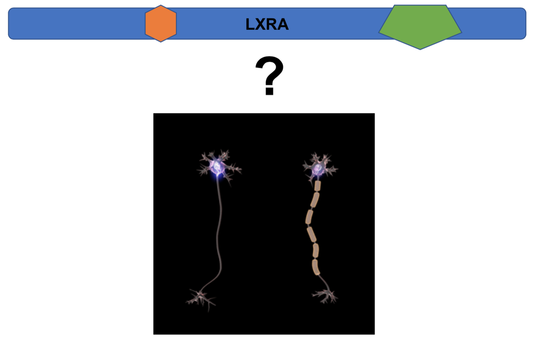
Myelin sheaths wrap around nerve axons and are composed mostly of lipids. LXRA is interesting to study in regards to MS due to the fact that it plays a role in lipid homeostasis, inflammation, and immune response. LXRA has two domain regions with one being a zinc finger domain that can bid to DNA and the other being a ligand binding domain. For this study I wanted to focus on the ligand binding domain. One paper found that 70% of patients in their study had the same mutation in this domain that changed an arginine to a glutamine [3]. LXRA is highly conserved throughout model organisms. The model organisms that will be used are mice and zebrafish. They both are very good models for studying myelination and have high conservation to humans. There is a mouse model for studying demyelination disease called EAE mice. Zebrafish can be used for high-throughput assays and are transparent so they allow for easy assays to view myelin sheaths. The overall goal of this study is to determine the role of LXRA in myelination.
Aim 1
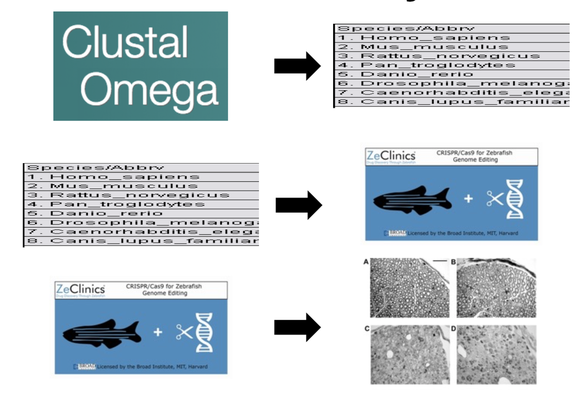
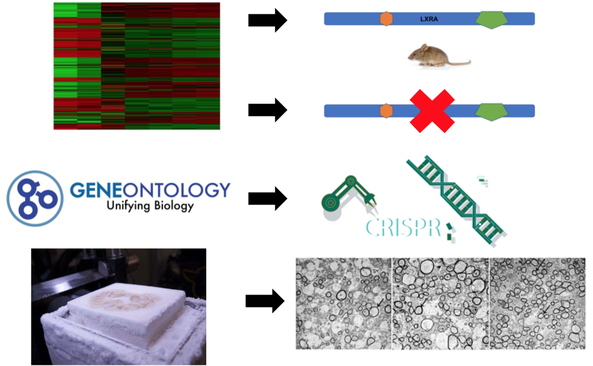
The first part of my study will be to identify the importantly conserved amino acids in the ligand binding domain of the LXRA protein. To do this I will use ClustalOmega to align sequences of different organisms. From there I will look for the most conserved amino acids among the organisms. I will then use CRISPR/Cas9 to remove these amino acids one at a time in zebrafish. Then an Osmium Tetroxide assay that will stain for myelin to determine the difference in myelination of mutant and wild type zebrafish. By performing this part of the study I think I will determine that mutations made to the ligand binding domain of the LXRA protein will lead to a reduction in myelination.
Aim 2
The second aim of my study will try to characterize differentially expressed genes in the central nervous system in LXRA deficient mice. To do this, I will perform RNA-seq in mice with LXRA and mice deficient of LXRA. I will then sort these genes using Gene Ontology. From here I will use CRISPR/Cas9 to determine the genes that are important for myelination by sectioning brains and spinal cords of mice and staining them with Osmium Tetroxide to look for myelination. I believe this study will show that genes involved in lipid homeostasis will be down-regulated in mutant mice and lead to demyelination.
Aim 3
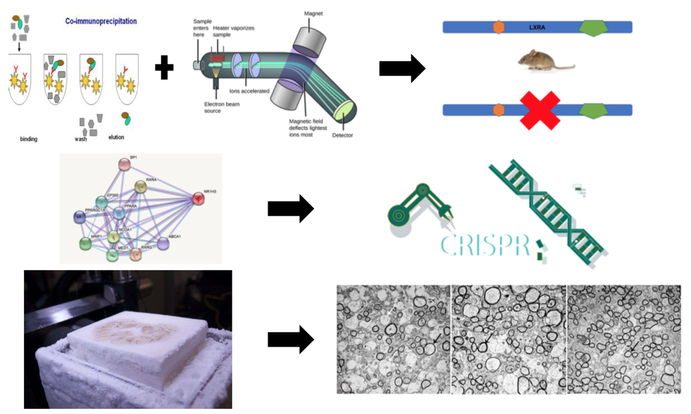
The third aim will be to determine the different protein interactions between WT and mutant LXRA mice. Co-IP and mass-spec will be performed from WT mice and mutant LXRA mice. From there I will mutant the proteins that differ using CRISPR/Cas9 in WT mice to determine which proteins are crucial for myelination. Brain and spinal cords will be sectioned and stained with Osmium Tetroxide to look at myelination levels. I believe this study will show that proteins involved in lipid homeostasis with differ between WT and mutant mice and that these proteins when mutant lead to lower myelination levels.
Future Directions
After conducting these experiments, I would like to determine if there are any possible drug therapy treatments that can be developed to target LXRA. I would also like to look more into how mutated LXRA may play a role in the vitamin D deficiency seen in many MS patients and whether this could be another pathway important in myelination.
| richardspresentationfinal5_1_182.pdf | |
| File Size: | 8342 kb |
| File Type: | |
| richardspresentationfinal5_1_18.pptx | |
| File Size: | 4679 kb |
| File Type: | pptx |
| richardspresentationdraft4_5_18.pptx | |
| File Size: | 596 kb |
| File Type: | pptx |
| richardspresentationdraft4_12_18.pptx | |
| File Size: | 1489 kb |
| File Type: | pptx |
References:
[1] Multiple sclerosis. (2017, August 04). Retrieved February 01, 2018, from https://www.mayoclinic.org/diseases-conditions/multiple-sclerosis/symptoms-causes/syc-20350269
[2] Definition of MS. (n.d.). Retrieved February 01, 2018, from https://www.nationalmssociety.org/What-is-MS/Definition-of-MS
[3] Wang, Z., Sadovnick, A. D., Traboulsee, A. L., Ross, J. P., Bernales, C. Q., Encarnacion, M., … Vilariño-Güell, C. (2016). Nuclear receptor NR1H3 in familial multiple sclerosis. Neuron, 90(5), 948–954. http://doi.org/10.1016/j.neuron.2016.04.039
[2] Definition of MS. (n.d.). Retrieved February 01, 2018, from https://www.nationalmssociety.org/What-is-MS/Definition-of-MS
[3] Wang, Z., Sadovnick, A. D., Traboulsee, A. L., Ross, J. P., Bernales, C. Q., Encarnacion, M., … Vilariño-Güell, C. (2016). Nuclear receptor NR1H3 in familial multiple sclerosis. Neuron, 90(5), 948–954. http://doi.org/10.1016/j.neuron.2016.04.039