This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is a protein interaction network?
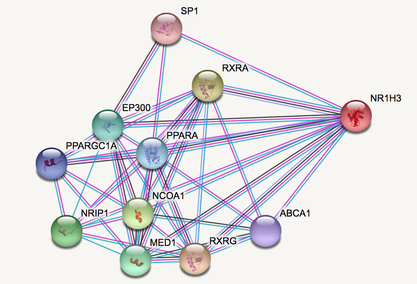
Throughout the cell, proteins interact with one another. These interactions are mapped with protein interaction networks. Interaction networks can be very important to determine how proteins are working together to accomplish a goal. They can also be used to determine what proteins active or repress another protein [1]. LXRA has been shown to form heterodimers with another protein, RXR [2]. It is important to study how LXRA can interact with other proteins to determine the role it plays in myelination.
How to generate a protein interaction network.
The database STRING allows us to easily create interaction networks. All we have to do is put in or protein of choice into the search and it will automatically generate a network that the protein directly interacts with. It can also be "zoomed out" to see secondary interactions.
Discussion
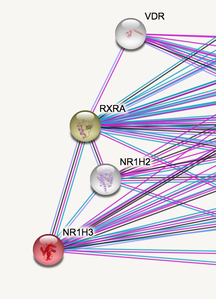
After creating protein interaction networks for LXRA, I was able to see a connection with a gene that is a receptor for Vitamin D (VDR). It has been shown that many patients with MS have Vitamin D deficiency [3]. This is a very interesting discovery because the LXRA gene may play role in how Vitamin D is taken up by the body. This may be a good direction to go in order to determine of LXRA plays a role in the MS disease.
References:
[Header Image] SC13 Research Highlight: Large Graph Processing Without the Overhead. (2013, November 20). Retrieved April 13, 2018, from https://www.hpcwire.com/2013/11/16/sc13-research-highlight-large-graph-processing-without-overhead/#foobox-1/0/image2.jpg
[1] Vazquez, A., & Alzate, O. (n.d.). Protein Interaction Networks. Retrieved April 13, 2018, from https://www.ncbi.nlm.nih.gov/pubmed/21882453
[2] Wang, Z., Sadovnick, A. D., Traboulsee, A. L., Ross, J. P., Bernales, C. Q., Encarnacion, M., … Vilariño-Güell, C. (2016). Nuclear receptor NR1H3 in familial multiple sclerosis. Neuron, 90(5), 948–954. http://doi.org/10.1016/j.neuron.2016.04.039
[3] Elkama, A., & Karahalil, B. (2018). Role of gene polymorphisms in vitamin D metabolism and in multiple sclerosis. Archives of Industrial Hygiene and Toxicology, 69(1), 25-31. doi:10.2478/aiht-2018-69-3065
[Header Image] SC13 Research Highlight: Large Graph Processing Without the Overhead. (2013, November 20). Retrieved April 13, 2018, from https://www.hpcwire.com/2013/11/16/sc13-research-highlight-large-graph-processing-without-overhead/#foobox-1/0/image2.jpg
[1] Vazquez, A., & Alzate, O. (n.d.). Protein Interaction Networks. Retrieved April 13, 2018, from https://www.ncbi.nlm.nih.gov/pubmed/21882453
[2] Wang, Z., Sadovnick, A. D., Traboulsee, A. L., Ross, J. P., Bernales, C. Q., Encarnacion, M., … Vilariño-Güell, C. (2016). Nuclear receptor NR1H3 in familial multiple sclerosis. Neuron, 90(5), 948–954. http://doi.org/10.1016/j.neuron.2016.04.039
[3] Elkama, A., & Karahalil, B. (2018). Role of gene polymorphisms in vitamin D metabolism and in multiple sclerosis. Archives of Industrial Hygiene and Toxicology, 69(1), 25-31. doi:10.2478/aiht-2018-69-3065