This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are post-translational modifications?
There are many post-translation modifications. They all, in some way, increase functional diversity of the proteome. These modifications can cleave subunits, degrade proteins, or add chemical groups onto parts of the protein. This is done through methylation, phosphorylation, ubiquitination, proteolysis, and others. The modifications play roles in normal and mutant proteins [1]. This is why studying post-translational modifications is very important to better understand disease pathogenesis.
How to determine phosphorylation sites.
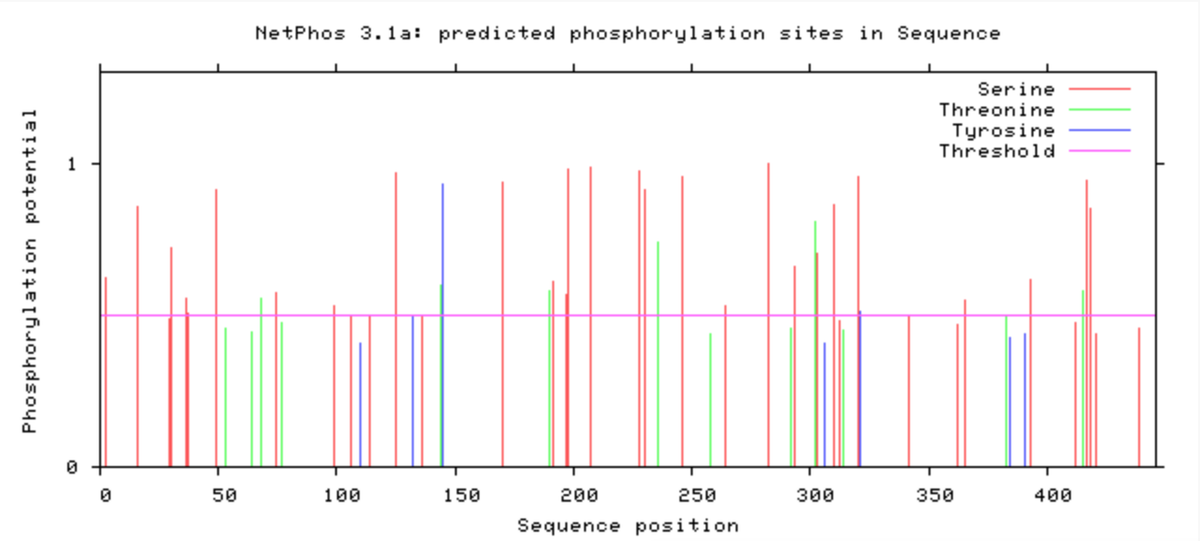
The database NetPhos, among others, can provide potential phosphorylation sites for a given protein. It is likely that the protein would require a serine, threonine, or tyrosine amino acid residue. The database uses these sites to predict where phosphorylation can occur. The following image shows phosphorylation sites for the LXRA protein (Fig. 1).
Discussion
Determining the possible phosphorylation sites of proteins can be key in determining how proteins work and how they interact with other proteins. It can give researchers a better idea of how proteins work normally and mutated. NetPhos showed that LXRA was phosphorylated at the 415 amino acid site by Protein Kinase C (PKC). This is particularly interesting considering it was determined that a common amino acid mutation that changes and arginine to a glutamine at this position [2].
References:
[Header Image] Post Translational Modifications: An Overview. (n.d.). Retrieved April 13, 2018, from https://www.ptglab.com/news/blog/post-translational-modifications-an-overview/
[1] Overview of Post-Translational Modifications (PTMs). (n.d.). Retrieved April 13, 2018, from https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
[3] Wang, Z., Sadovnick, A. D., Traboulsee, A. L., Ross, J. P., Bernales, C. Q., Encarnacion, M., … Vilariño-Güell, C. (2016). Nuclear receptor NR1H3 in familial multiple sclerosis. Neuron, 90(5), 948–954. http://doi.org/10.1016/j.neuron.2016.04.039
[Header Image] Post Translational Modifications: An Overview. (n.d.). Retrieved April 13, 2018, from https://www.ptglab.com/news/blog/post-translational-modifications-an-overview/
[1] Overview of Post-Translational Modifications (PTMs). (n.d.). Retrieved April 13, 2018, from https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
[3] Wang, Z., Sadovnick, A. D., Traboulsee, A. L., Ross, J. P., Bernales, C. Q., Encarnacion, M., … Vilariño-Güell, C. (2016). Nuclear receptor NR1H3 in familial multiple sclerosis. Neuron, 90(5), 948–954. http://doi.org/10.1016/j.neuron.2016.04.039